Research Outcomes
This section presents the main achievements and research outcomes obtained during the project timeline. Our work has advanced the state-of-the-art in medical AI through several key contributions.
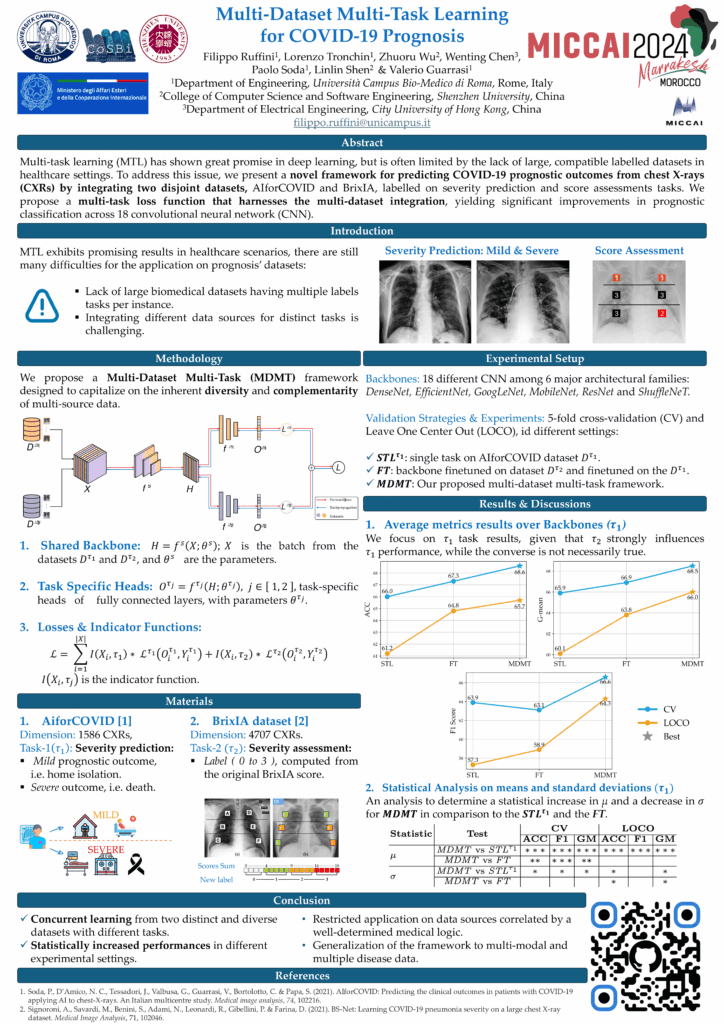
Multi-Dataset Multi-Task Learning for COVID-19 Prognosis
In the fight against the COVID-19 pandemic, leveraging artificial intelligence to predict disease outcomes from chest radiographic images represents a significant scientific aim. The challenge, however, lies in the scarcity of large, labeled datasets with compatible tasks for training deep learning models without leading to overfitting. Addressing this issue, we introduce a novel multi-dataset multi-task training framework that predicts COVID-19 prognostic outcomes from chest X-rays (CXR) by integrating correlated datasets from disparate sources. Our framework hypothesizes that assessing severity scores enhances the model's ability to classify prognostic severity groups, thereby improving its robustness and predictive power. The proposed architecture integrates two publicly available CXR datasets, AIforCOVID for severity prognostic prediction and BRIXIA for severity score assessment, demonstrating significant performance improvements across 18 different neural network architectures.
Recognition: This research was presented at the 27th International Conference on Medical Image Computing and Computer Assisted Intervention (MICCAI 2024), held in Marrakech, Morocco (October 6-10, 2024).
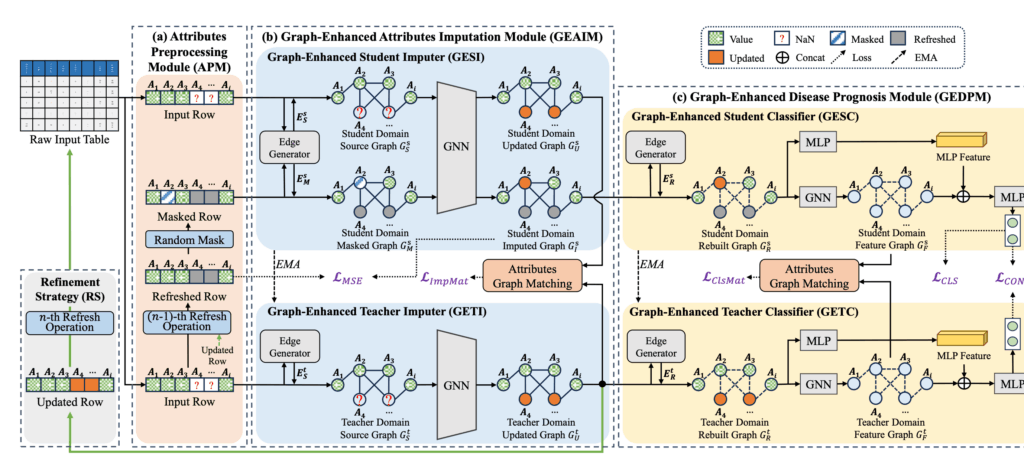
Handling Missing Clinical Data with Graph-Based AI
Clinical datasets often suffer from missing values due to the hectic conditions of hospital workflows, especially during pandemic outbreaks. To address this challenge, we developed ACGM (Attribute-Centric Graph Modeling), an innovative framework that simultaneously handles missing data imputation and disease prognosis. The key insight is to model clinical attributes as nodes in a graph, where relationships between different medical measurements (such as blood markers, vital signs, and symptoms) are explicitly captured. This approach allows the AI to leverage the inherent correlations among clinical variables to intelligently fill in missing values while predicting patient outcomes. Our interpretability analysis revealed that attributes such as LDH (Lactate Dehydrogenase), breathing difficulty, and oxygen saturation (SaO2) significantly impact COVID-19 prognosis, aligning with clinical evidence.
Recognition: This work has been accepted for publication in IEEE Journal of Biomedical and Health Informatics.
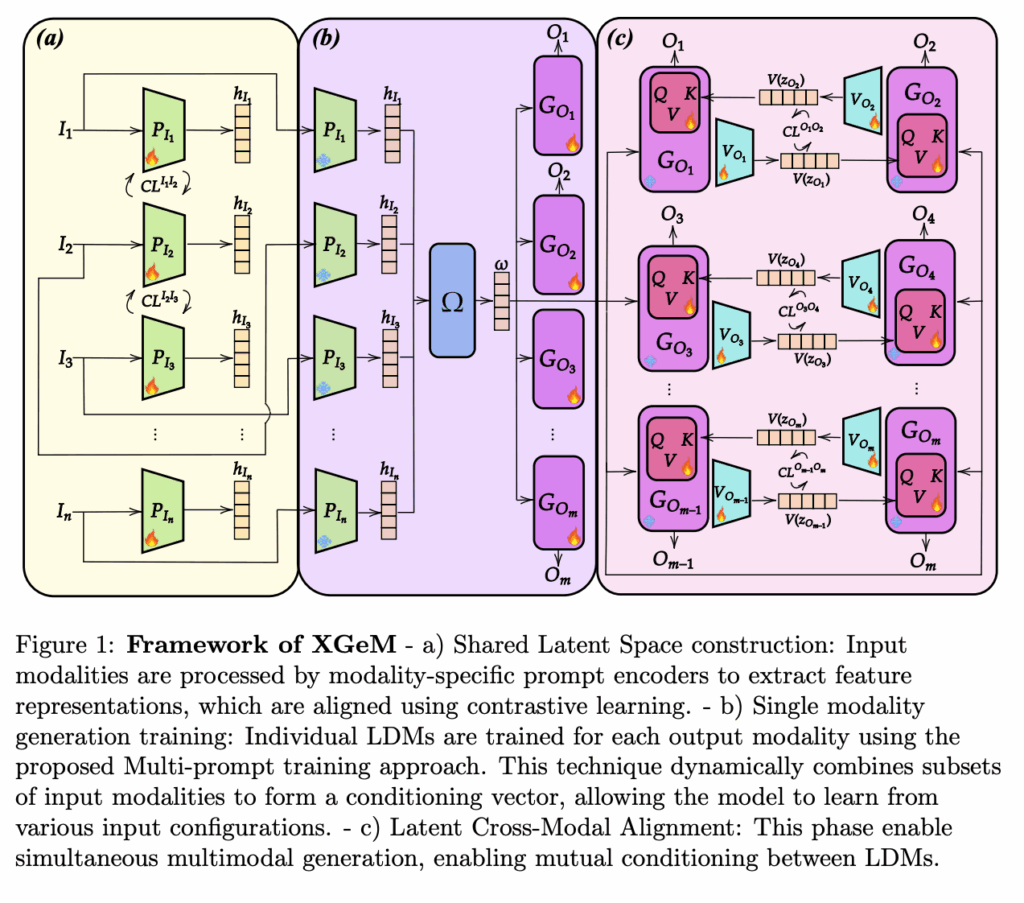
Synthetic Medical Data Generation for AI Training
One of the major bottlenecks in medical AI is the scarcity of annotated clinical data due to privacy concerns and the cost of expert annotations. To overcome this limitation, we developed XGeM, a foundation model capable of generating realistic synthetic medical data across multiple modalities. The model can generate chest X-rays (both frontal and lateral views) along with corresponding radiological reports, creating coherent multimodal synthetic samples. This "any-to-any" generation capability means the model can produce any combination of outputs from any input: generating a report from an image, an image from text, or completing missing views. Clinical validation through Turing tests with expert radiologists confirmed the high quality and clinical plausibility of the generated data. A public demo is available at: https://xgem.ucbm.org/real-demo
Recognition: This research has been submitted to Computerized Medical Imaging and Graphics (Elsevier) and presented at International Joint Conference on Neural Networks (IJCNN).
Code available at: https://github.com/cosbidev/XGeM
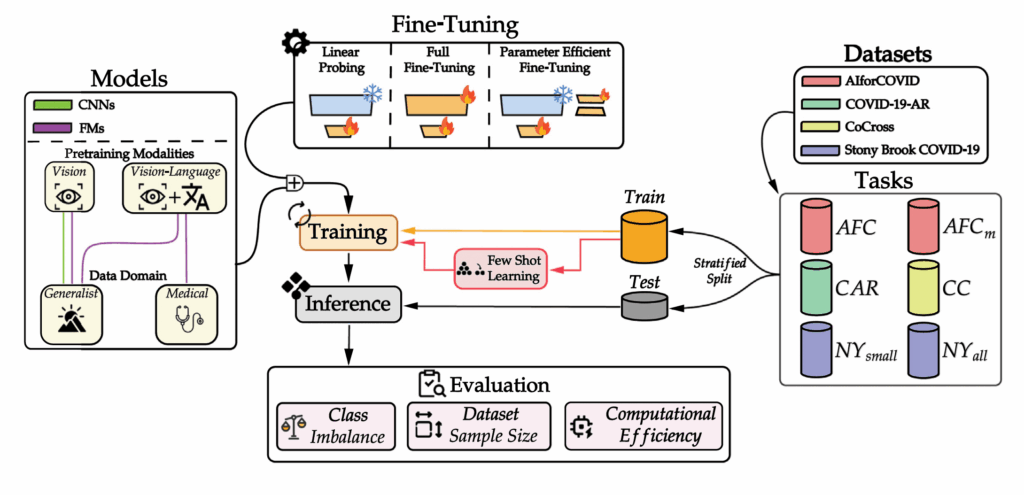
Benchmarking AI Models for COVID-19 Prognosis
Understanding which AI models work best under different clinical conditions is crucial for real-world deployment. We conducted the first comprehensive benchmark comparing traditional convolutional neural networks (CNNs) with modern Foundation Models for predicting COVID-19 patient outcomes from chest X-rays. Our study evaluated multiple adaptation strategies, including parameter-efficient fine-tuning methods that update only a small fraction of model parameters. Key findings show that traditional CNNs remain reliable when data is scarce or highly imbalanced, while Foundation Models excel when sufficient training data is available. These results provide practical guidance for clinicians and AI developers in selecting the most appropriate models based on their specific data constraints.
Recognition: Published in Computer Methods and Programs in Biomedicine (Elsevier, 2026). Code available at: https://github.com/cosbidev/PEFT_Prognosis
AI-Powered Tumor Segmentation
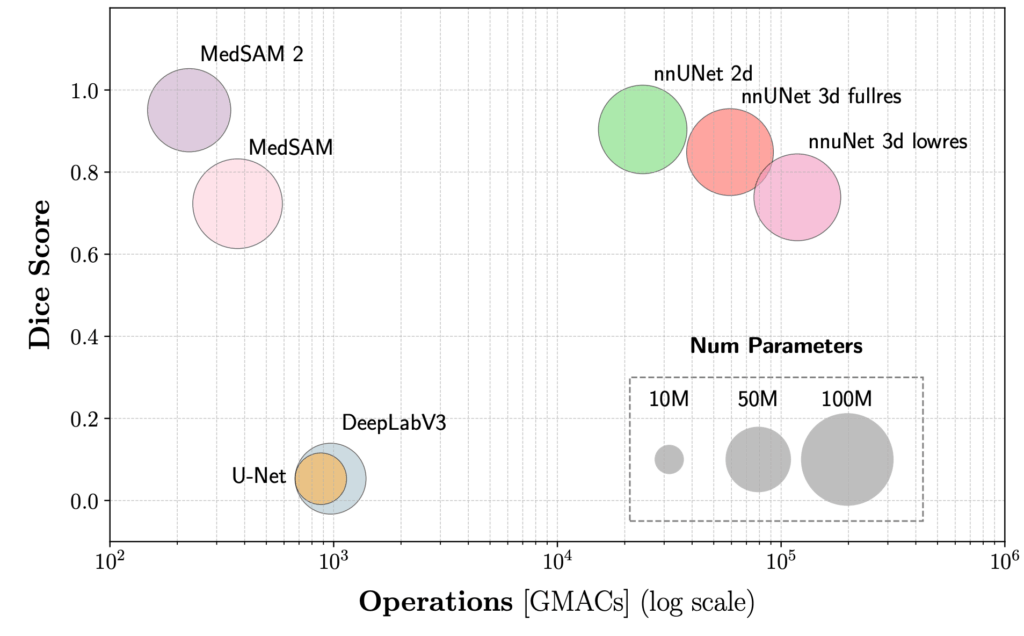
Accurate delineation of tumors in medical images is essential for diagnosis and treatment planning. We explored two complementary directions in this area. First, we conducted a systematic benchmark comparing traditional segmentation models (U-Net, DeepLabV3) with modern foundation models (MedSAM, MedSAM 2) for lung tumor segmentation in CT images. Results showed that foundation models, particularly MedSAM 2, achieve superior accuracy with lower computational costs. Second, we developed vMambaX, a lightweight framework for multimodal PET-CT tumor segmentation. By intelligently combining anatomical information from CT scans with metabolic information from PET imaging through an adaptive gating mechanism, vMambaX achieves state-of-the-art segmentation performance while maintaining computational efficiency suitable for clinical workflows.
Recognition: The benchmarking study was published at IEEE CBMS 2025. The vMambaX framework has been submitted to IEEE ISBI 2025.
Publications
Journal Articles
- Wu, Z., Chen, W., Li, X., Ruffini, F., et al. (2025). "ACGM: Attribute-Centric Graph Modeling Network for Concurrent Missing Tabular Data Imputation and COVID-19 Prognosis". IEEE Journal of Biomedical and Health Informatics. DOI: 10.1109/JBHI.2025.3618935
- Ruffini, F., Mulero Ayllón, E., Shen, L., Soda, P., and Guarrasi, V. (2026). "Benchmarking foundation models and parameter-efficient fine-tuning for prognosis prediction in medical imaging". Computer Methods and Programs in Biomedicine, 275, 109196.
- Molino, D., et al. (2025). "MedCoDi-M: A Multi-Prompt Foundation Model for Multimodal Medical Data Generation". Submitted to Information Fusion.
Conference Papers
- Ruffini, F., et al. (2024). "Multi-Dataset Multi-Task Learning for COVID-19 Prognosis". MICCAI 2024, pp. 251-261.
- Mulero Ayllón, E., et al. (2025). "Can Foundation Models Really Segment Tumors? A Benchmarking Odyssey in Lung CT Imaging". IEEE CBMS 2025, pp. 375-380.
- Molino, Daniele, Francesco Di Feola, Eliodoro Faiella, Deborah Fazzini, Domiziana Santucci, Linlin Shen, Valerio Guarrasi, and Paolo Soda. "MedCoDi-M: A Multi-Prompt Foundation Model for Multimodal Medical Data Generation." arXiv preprint arXiv:2501.04614 (2025).
- Mulero Ayllón, E., et al. (2025). "Context-Gated Cross-Modal Perception with Visual Mamba for PET-CT Lung Tumor Segmentation". Accepted to IEEE ISBI 2026.
- Aksu, F., et al. (2024). "Towards AI-driven next generation personalized healthcare and well-being". Ital-IA 2024, pp. 360-365.